|
http://www.abbs.info e-mail:[email protected] ISSN
0582-9879
ACTA BIOCHIMICA
et BIOPHYSICA SINICA 2003, 35(11): 1029-1034
CN 31-1300/Q |
Short Communication
Detection
and Genotyping of Human Papillomavirus DNA in Cervical
Cancer Tissues with Fluorescence Polarization
GAO Yan-E*, ZHANG Ju1, WU Jing,
CHEN Zhong-Can1, YAN Xiao-Jun1
( Department of Obstetrics and Gynecology, Second Hospital, Xi’an Jiaotong
University, Xi’an 710004, China;
1Institute of Gene Diagnosis, Fourth Military Medical University, Xi’an
710032, China )
Abstract To evaluate the type-specific prevalence of eight common types of human papillomavirus (HPV) in patients with cervical cancer living in Shanxi, China, with fluorescence polarization detection, crude DNA extracted from 137 samples of early-stage cervical cancer (within stage IIa) and chronic cervicitis was subjected to HPV L1 consensus GP5+/GP6+ system. Then, the HPV-positive products identified by GP5+/GP6+ PCR were genotyped based on template-directed dye-terminator incorporation assay with fluorescence polarization detection (TDI-FP): the PCR products were respectively hybridized with designed type-specific probes within the GP5+/GP6+ amplicons for eight common HPV types (HPV 6, 11, 18, 16, 31, 33, 35, and 58), and specific fluorescence-labeled ddNTPs (TAMRA-ddTTP or R110-ddGTP) were directly incorporated to the ends of the corresponding probes under directing of the corresponding template in PCR products, which was reflected and read by high FP values for TAMRA or R110. HPV DNA was detected in 38.89% (28/72) cases of chronic cervicitis, and 87.69% (57/65) cases of cervical cancer. There was a significant difference in HPV prevalence between these two groups. The four commonly identified types in patients with cervical cancer were HPV 16 (45.6%), HPV 18 (22.8%), HPV 58 (17.5%), and HPV 31 (7.02%), and in those with chronic cervicitis were HPV 16 (35.7%), HPV 11 (32.1%), HPV 6 (21.4%), and HPV 18 (10.7%). 57.14% of HPV types detected in patients with chronic cervicitis were high-risk types. HPV 16 was the most common viral type identified in both groups. Type-specific prevalence of HPV DNA has some characteristics in patients with chronic cerviticis and cervical cancer living in Shanxi, China,and the worldwide uncommon type HPV 58 is relatively common in both kinds of cases. The high prevalence of HPV 58 in Chinese women should been considered in diagnosis and vaccine designs of HPV.
Key words cervical cancer; human papillomavirus (HPV); genotyping; fluorescence polarization
Human papillomavirus
(HPV) is one of the most common causes of sexually transmitted diseases
in both men and women worldwide. To date, more than 100 distinct human papillomavirus
genotypes have been described, and near half of them can infect the genital
tract which can be classified into high-risk and low-risk groups, as established
by their associated clinical prevalence. The HPV genotypes 6, 11, 16, 18,
31, 33, 35 and 58 have been detected most frequently in mucosal lesions.
Among them, HPV 16, 18, 31, 33, 35 and 58 have been defined as high-risk
genotypes based on their prevalence in cervical intraepithelial neoplasia
(CIN) lesions and cervical cancer.
Cervical cancer is preceded only by breast cancer as the most common cause
of death from cancer in women worldwide. In developing countries, cervical
cancer is the most common cancer in women and may constitute up to 25% of
all female cancers[1]. Since 1980s, the link between HPV and cervical cancer
has been well established. HPV has been implicated in 99.7% of cervical
cancer worldwide[2]. In 1996, the World Health Organization, along with
the European Research Organization on Genital Infection and Neoplasia and
the National Institutes of Health Consensus Conference on Cervical Cancer,
recognized HPV as an important cause of cervical cancer[3].
Cervical cancer is also relatively common in China, with the occurrence
of 150 000 new cases each year, and certain parts of China, such as Luoyang
county in Shanxi province, have the highest incidence of cervical cancer
in the world[4,5]. Although HPV type 16 is the prevailing high-risk type,
being present in about 50% of the cervical cancer worldwide[2, 6-8], geographic
variation has been observed in some less common HPV types in different areas
of the world[9]. Understanding the area-specific distribution of high-risk
HPV types has important implications for the future studies on the diagnostic
approach and prophylactic and therapeutic vaccine designs of cervical cancer.
To elucidate further the epidemiology of HPV infection in the Chinese population,
we sought to determine the type-specific prevalence of the most common HPVs
including HPV 6, 11, 16, 18, 31, 33, 35 and 58 in women with cervical lesions
living in Shanxi Province, China, using the novel HPV typing method based
on template-directed dye-terminator incorporation assay with fluorescence
polarization detection (TDI-FP) we developed previously[10].
1 Materials and
Methods
1.1 Materials
Fluorescence polarization-capable instrument-victor2, shrimp alkaline phosphatase,
E. coli exonuclease I, AmpliTag DNA Polymerase, mixture of TAMRA-ddTTP and
R110-ddGTP were purchased from PerkinElmer (USA). 384-well blank-skirted
plates were obtained from MJ Research (USA). DNA marker (DL2000) was from
TaKaRa (Dalian, China). All primers and nucleotide probes were synthesized
in Sbsbio (Beijing, China).
1.2 Patients
Between September 2001 and March 2003, women visiting the Department
of Gynecology at the Second Hospital of Xi’an Jiaotong University for further
management following abnormal pap smears or cervical cancer were included
in the study. Biopsy tissue samples were obtained under colposcopy. The
biopsy specimens were divided into two parts, one of which was fixed in
10% buffered formalin for routine pathological diagnosis, the other was
suspended in 1 mL sterile 0.9% NaCl and stored at -20 ℃ for HPV analysis.
The diagnosis of cervical lesions was based on pathological findings. A
written informed consent was obtained from all participants and the study
was approved by the ethics committee of the hospital.
1.3 DNA extraction
Each biopsy specimen was resuspended in 0.3 mL PBS, repeatedly frozen,
thawed, and minced. The suspension was mixed with 0.5 mL of proteinase K
(200 g/L) in TE buffer (10 mmol/L Tris, 1 mmol/L EDTA) and incubated at
56 ℃ for 5 h. Then, the mixture was boiled for 10 min for inactivating proteinase
K, chilled on ice, and centrifuged at 12 000 g for 5 min. The supernatant
was decanted and stored at -20 ℃ until amplification. 2 μL of the supernatant
was used directly for each PCR analysis.
1.4 HPV detection using GP5+/GP6+ consensus primer set
All the samples were tested for HPV processed a conventional PCR with
GP5+ and GP6+ primers as described previously[11] with some modification.
In brief, each GP5+/GP6+ PCR reaction was carried out in 50 μL containing
150 μmol/L dNTP, 3.0 mmol/L MgCl2, 0.5 u of Taq DNA polymerase, 0.25 μmol/L
each of the GP5+ [5′-TTTGTTACTGTG(T)GTA(G)-GATACT(C)AC-3′] and GP6+ [3′-CTT(C)AT(A)-ACTAAATGTC(T)AAAT(C)AA(C)AAAG-5′],
and 2 μL of the processed supernatant. A 3-min denaturation step at 94 ℃
was followed by 40 cycles of amplification in a thermal cycler. Each cycle
included a denaturation step at 94 ℃ for 1 min, a primer annealing step
at 45 ℃ for 1 min, and a chain elongation step at 72 ℃ for 2 min. The final
elongation step was prolonged by 5 min to ensure a complete extension of
the amplified DNA. For confirming the accuracy of HPV typing assays, sequence-verified
controls for each HPV genotype and blank control were included in each assay
from this step.
The fragments amplified by GP5+/GP6+ PCR were 150 bp locating in the L1
regions of HPVs. The samples from which 150 bp fragments were amplified
were identified as HPV-positive. For HPV-negative samples determined by
GP5+/GP6+ PCR, the DNA quality of the samples was assessed by PCR amplification
of the β-globin gene using primers PC04 (5′-CAACTTCATCCACGTTCACC-3′) and
PH20 (5′-GAAGAGCCAAGGACAGGTAC-3′), resulting in a 268 bp product[12], to
confirm the presence of an adequate preparation of DNA and that nonspecific
inhibitors were absent.
Each PCR product was analysis by electro-phoresis on 2.0% agarose gels stained
with ethidium bromide.
1.5 HPV typing with TDI-FP assay
Type-specific probes for the eight common HPV types were designed by
DNA Star within the GP5+/GP6+ amplification regions of the L1 genes. The
sequences of the probes are given in Table 1. The GP5+/GP6+ PCR products
of HPV-positive samples were typed by type-specific probes based on TDI-FP.
In order to eliminate the excess consensus GP5+/GP6+ primers and dNTPs used
in the PCR reaction, every 1 μL of PCR product was diluted by adding 9 μL
of ddH2O, and enzymatically digested by mixing up with shrimp alkaline phosphatase
(1 u), and E. coli exonuclease I (1 u) in shrimp alkaline phosphatase buffer
(0.5 mol/L Tris・HCl, pH 8.5, 50 mmol/L MgCl2) at 37 ℃ for 2 h. The enzymes
were inactivated by heating at 95 ℃ for 20 min.
7 μL of the enzymatically pretreated GP5+/GP6+ PCR product was mixed with
13 μL of TDI-FP mixture containing 10× reaction buffer, AmpliTaq DNA polymerase,
TAMRA-ddTTP/R110-ddGTP, and individual TDI primers of eight HPV types, i.e.,
type-specific oligonucleotide probes for HPV 6, 11, 16, 18, 31, 33, 35 or
58, and denatured at 95 ℃ for 2 min, followed by 30 cycles of 95 ℃ for 15
s and 50 ℃ for 30 s. At the end of cycles, the mixture was held at 4 ℃.
The fluorescence polarization (FP) values for TAMRA and R110 were measured
and analyzed on Fluorescence polarization-capable instrument-victor2.
The HPV type was determined by corresponding probe hybridization followed
by template-directed single-dye-base probe extension, i.e., template-directed
dye-terminator incorporation specifically to the probe, which was recognized
by high FP value for TAMRA or R110. The dye-terminators, TAMRA-ddTTP and
R110-ddGTP designed for the corresponding HPV types are shown in Table 1.
Table 1 HPV type-specific probes within GP5+/GP6+ amplicons and their corresponding dye-terminators
|
HPV type |
Probe sequence (5′ →3) | Dye-terminator |
| 6 | ATCCGTAACTACATCTTCCACATACACCA | TAMRA-ddTTP |
| 11 | ATCTGTGTCTAAATCTGCTACATACACTA | TAMRA-ddTTP |
| 16 | GTCATTATGTGCTGCCATATGTACTTCAG | TAMRA-ddTTP |
| 18 | TGCTTCTACACAGTCTCCTGTACCTGGGC | TAMRA-ddTTP |
| 31 | TGTTTGTGCTGCAATTGCAAACAGTGATA | R110-ddGTP |
| 33 | TTTATGCACACAAGTAACTAGTGACAGTA | R110-ddGTP |
| 35 | GTCTGTGTGTTCTGCTGTCTTCTAGTGA | R110-ddGTP |
| 58 | ATTATGCACTGAAAGTAACTAAGGAAGGTA | R110-ddGTP |
1.6 Statistical
analysis
χ2 test was used to compare the proportions of HPV infections among
women with chronic cervicitis and cervical cancer. Difference was regarded
as significant when P<0.05.
2 Results
2.1 Clinicopathological data
A total of 137 Chinese women who had cervical samples with an adequate
quality of DNA were included in the study. Among them, 72 had chronic cervicitis,
and 65 had cervical cancer. The average age of patients with chronic cervicitis
was 42 years (range from 25 to 56 years), and of those with cervical cancer
was 48 years (range from 28 to 71 years). According to the staging categories
of the International Federation in Gynecology and Obstetrics (FIGO), all
patients with cervical cancer were classified as staged I or IIa.
2.2 HPV DNA detection using GP5+/GP6+ consensus primers
Overall, 85 of the 137 samples were identified to be HPV-positive for
having 150 bp bands in 2.0% agarose gel electrophoresis after GP5+/GP6+
PCR (Fig. 1), with a prevalence of 38.89% (28/72) for chronic cervicitis,
and 87.69% (57/65) for cervical cancer. The difference in HPV prevalence
between these two groups was statistically significant (P<0.01).
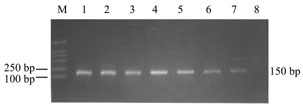
Fig.1 2.0% agarose
gel electrophoresis of GP5+/GP6+ PCR products of HPV DNA
M, DNA marker; 1-7, HPV-positive cervical specimens; 8, blank control.
2.3 HPV typing
with TDI-FP assay
GP5+/GP6+ PCR products from 85 HPV-positive samples were respectively
detected for the eight HPV types with TDI-FP assay. The results are plotted
and shown in Fig.2.
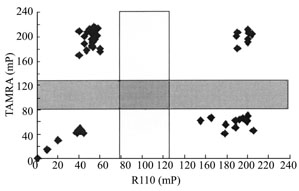
Fig.2 Scatter
plot result of HPV genotyping using TDI-FP
Blank controls, or samples infected with undetermined HPV types have
low FP values for both dyes, occupying the lower left corner. In the upper
left corner, the samples infected with HPV 6, 11, 16, or 18 have high FP
for TAMRA but low FP for R110. The samples occupying the lower right corner,
with high R110 but low TAMRA FP values, exhibit being infected with HPV
31, 33, 35, or 58. Samples found in the right upper corner with high FP
values for both dyes are infected with multiple corresponding HPV types.
Following a standard protocol, the FP values cluster into four groups. As expected, the blank controls without DNA, or samples infected with undetermined HPV types (HPV X) have low FP values for both dyes, due to no probe-hybridization and dye-terminators incorporation but the free remaining of the dye terminators in the solutions, occupying the lower left corner. In the upper left corner, the samples have high FP for TAMRA but low FP for R110, reflecting significant template-directed incorporation of the TAMRA-ddTTP terminator to the corresponding type-specific probes (HPV 6, 11, 16, or 18) but minimal incorporation of the R110-ddGTP as expected. On the other hand, the samples occupying the lower right corner, with high R110 but low TAMRA FP values, exhibit significant template-directed incorporation of the R110-ddGTP terminator to the corresponding type-specific probes (HPV 31, 33, 35, or 58) but minimal incorporation of the TAMRA-ddTTP. Samples found in the right upper corner with high FP values for both dyes indicate significant template-directed incorporation of both dye terminators to the corresponding type-specific probes, and these individuals are infected with multiple corresponding HPV types. FP values for R110 and TAMRA for each HPV type are presented in Table 2.
Table 2 FP values for R110 and TAMRA for each HPV type
| HPV type (n) | TAMRA* | R110* |
| 6(7) | 195.58±9.71 | 49.02±5.3811 |
| (10) | 205.02±8.31 | 48.80±5.3216 |
| (36) | 218.97±4.26 | 45.32±3.3218 |
| (16) | 192.89±6.27 | 47.56±5.8331 |
| (5) | 46.73±10.55 | 195.32±9.8633 |
| (2) | 50.66 | 206.1235 |
| (2) | 60.32 | 217.3258 |
| (12) | 56.04±8.12 | 186.57±7.16 |
| X (12) | 55.26±7.27 | 56.94±7.68 |
*Data are represented as x±s.
Table 3 HPV types identified in chronic cervicitis and cervical cancer
| Diagnosis |
HPV |
Low-risk types | High-risk types | Unknown types |
Multiple types |
||||||
| 6 | 11 | 16 | 18 | 31 | 33 | 35 | 58 | X | |||
| Chronic cervicitis |
28/72 |
6/28 (21.4) |
9/28 (32.1) |
10/28 (35.7) |
3/28 (10.7) |
1/28 (3.57) |
0/28 (0) |
0/28 (0) |
2/28 (7.14) |
4/28 (14.29) |
4/28 (14.29) |
| Cervical cancer | 57/65 (87.69) |
1/57 (1.75) |
1/57 (1.75) |
26/57 (45.6) |
13/57 (22.8) |
4/57 (7.02) |
2/57 |
2/57 (3.51) |
10/57 (17.5) |
8/57 (14.04) |
7/57 (12.28) |
+, HPV-positive samples. Unknown types, undetermined HPV types. Numbers in parentheses are percentages of HPV types. The percentages of all HPV types detected in the diseases may add to more than 100% because of multiple infections.
The results of HPV typing by TDI-FP analysis are summarized in Table 3. The types identified most frequently in chronic cervicitis were HPV 16 (35.7%), HPV 11 (32.1%), HPV 6 (21.4%), HPV 18 (10.7%), and HPV 58 (7.14%), and in cervical cancer were HPV 16 (45.6%), HPV 18 (22.8%), HPV 58 (17.5%), and HPV 31 (7.02%). HPV 16 was the most common viral type identified in both groups either as single-type infection or as multiple-type infection. HPV 58 was the third common HPV type identified from patients with cervical cancer preceded only by HPV 16 and HPV 18. Undetermined HPVs (X types) were present in 14.29% of the HPV-positive patients with chronic cervicitis, and in 14.04% of those with cervical cancer. Among the HPV-positive patients, multiple-type infection was respectively detected in 14.29% of those with chronic cervicitis, and in 12.28% of those with cervical cancer. The proportion of multiple HPV infection was not significantly different in these two groups (P>0.05).
3 Discussion
In the present report, based on detection and genotyping of HPV by TDI-FP
method, the prevalence rate of HPV infection among Shanxi Chinese patients
with cervical cancer of early stages (stage I or IIa) was found to be 87.69%.
High-risk HPVs were accounted for 96.39% of whole HPV types detected in
the HPV-positive patients, with HPV 16 being the most prevalent viral type.
These results are similar to the results of previously published studies
around world and in China[6, 13-15].
The prevalence rate of HPV in patients with chronic cervicitis was 38.89%,
significantly lower than that in those with cervical cancer. However, because
the women with chronic cervicitis enrolled in this study accepted the colposcopy
and biopsy analysis due to abnormal Pap smears, being at risk for HPV infection,
the prevalence rate might be higher than the women with general inflamed
cervices.
Decades of studies have confirmed that cervical infection by high-risk HPV
types is a precursor event to cervical cancer[3]. We found that 57.14% of
HPV types detected in patients with chronic cervicitis in this study were
high-risk types, indicating that detection of high-risk HPVs could greatly
facilitate the identification of women at risk for cervical cancer.
A further finding of our study is the relatively high prevalence of HPV
58 in women with both cervical lesions. In an international study, HPV 58
was found only in 2% cervical cancer specimen from Africa, North and South
America, Southeast Asia, and Europe[6]. However, among the HPV-positive
patients in our study, HPV 58 was detected in 7.14% of those with chronic
cervicitis, and in 17.5% of those with cervical cancer. In women with cervical
cancer, the prevalence of HPV 58 (17.5%) was the third only to HPV 16 (45.6%)
and HPV 18 (22.8%). A high prevalence of this worldwide uncommon HPV type
has also been reported among Chinese women living in Taiwan[16], Shanghai[17],
and Hong Kong[8], particularly in those with cervical cancer. The distinct
high proportion of HPV 58 infection in Chinese women should been considered
in the design of HPV detection methods and the development of vaccines directing
against HPV.
It is agreed that early detection and subsequent intervention of HPV infection
is important in prevention, warning, and prognosis of cervical cancer[18].
As HPV cannot be cultured in the laboratory from clinical specimens and
immunologic assays are still not well established, the diagnosis of HPV
infections presently depends on molecular methods for detecting HPV DNA
sequences in clinical specimens.
Previous studies show that TDI-FP assay is simple, accurate, sensitive,
specific, and suitable for automated genotyping of large number of samples,
and has been used in large-scale analysis of single nucleotide polymorphisms
(SNPs)[19-21]. The novel method of HPV detection and genotyping used in
our study is based on TDI-FP assay. In this method, the sample is firstly
amplified by GP5+/GP6+ consensus primer set. Then in the presence of DNA
polymerase and fluorescence-labeled dNTPs (i.e., dye-labeled terminator),
a type-specific probe is annealed to the DNA fragment of corresponding type
within the PCR amplicon, and extended specifically by one dye-labeled terminator
directed by the template of the target DNA. Incorporation of a fluorescent
terminator into a oligonucleotide probe increases 10-fold molecular weight
of the fluorophore[19]. Since FP value is directly proportional to the molecular
weight of the fluorophore, a dye-labeled terminator attached to a probe
shows a higher FP value than a free dye-labeled terminator[19, 20]. The
FP values can be read by Fluorescence Polarization-Capable Instrument-Victor2.
The dNTP incorporated and corresponding HPV genotype can then be inferred.
In our study, we confirmed HPV genotyping results by including sequence-confirmed
controls of each HPV genotype and blank controls in each assay. 100% concordance
comparing TDI-FP with other methodologies has also been found in our previous
study[10] and been reported by others[20-22]. This HPV genotyping method
can detect eight HPV types in one-step PCR, and is high throughput and requires
no centrifugation, washing, separation, or transfer steps, resulting in
minimum hands-on time. So, it is suitable for large-scale HPV testing clinically.
References
1 Harro CD, Pang YY, Roden RB, Hildesheim A, Wang Z, Reynolds MJ, Mast
TC et al. Safety and immunogenicity trial in adult volunteers of a human
papillomavirus 16 L1 virus-like particle vaccine. J Natl Cancer Inst, 2001,
93: 284-292
2 Walboomers JM, Jacobs MV, Manos MM, Bosch FX, Kummer JA, Shah KV, Snijders
PJ et al. Human papillomavirus is a necessary cause of invasive cervical
cancer worldwide. J Pathol, 1999, 189(1): 12-19
3 Burd EM. Human papillomavirus and cervical cancer. Clin Microbiol Rev,
2003, 16(1): 1-17
4 Yang DW, Yao JF, Xing SF, Lin YX. Mass cytologic screening for cervical
carcinoma in China. A review of 7,735,057 reported cases. Acta Cytol, 1985,
29: 341-344
5 Zhang JM, Ruan SX,Liu DK, Gong HX, Yu BL, Bian SY, Zhang RT. The distribution
and characteristics of carcinoma of the cervix uteri in Lue Yang County.
Chin J Epidemiol, 1986, 7: 343-345
6 Bosch FX, Manos MM, Munoz N, Sherman M, Jansen AM, Peto J, Schiffman MH
et al. Prevalence of human papillomavirus in cervical cancer: a worldwide
perspective. International biological study on cervical cancer (IBSCC) Study
Group. J Natl Cancer Inst, 1995, 87(11): 796-802
7 Li J, liu BY, zur Hausen H, Wang H, Yang HZ, Li L, Liu W et al. Investigation
of human papillomavirus (HPV) infection in cervical carcinoma tissues in
Chinese women. Chin J Exp Clin Virol, 1996, 10: 50-55
8 Chan PK, Li WH, Chan MY, Ma WL, Cheung JL, Cheng AF. High prevalence of
human papillomavirus type 58 in Chinese women with cervical cancer and precancerous
lesions. J Med Virol, 1999, 59(2): 232-238
9 Koutsky L. Epidemiology of genital human papillomavirus infection. Am
J Med, 1997, 102: 3-8
10 Zhang J, Gao YE, Yan XJ, Yin GW, Bai YJ, Li D. Development of a high
throughput gene diagnosis assay for genotyping human papillomavirus. Chin
J Lab Med, 2003, 26(3): 1-3
11 de Roda Husman AM, Walboomers JM, van den Brule AJ, Meijer CJ, Snijders
PJ. The use of general primers GP5 and GP6 elongated at their 3’ ends with
adjacent highly conserved sequences improves human papillomavirus detection
by polymerase chain reaction. J Gen Virol, 1995, 76: 1057-1062
12 Bell DA, Taylor JA, Paulson DF, Bobertson CN, Mohler JL, Lucier GW. Genetic
risk and carcinogen exposure: A common inherited defect of the carcinogen-metabolism
gene glutathione S-tranferase M1 (GSTM1) that increases susceptibility to
bladder cancer. J Natl Cancer Inst, 1993, 85: 1159-1164
13 Jastreboff AM, Cymet T. Role of the human papilloma virus in the development
of cervical intraepithelial neoplasia and malignancy. Postgrad Med J, 2002,
78: 225-228
14 Munoz N, Bosch FX, de Sanjose S, Herrero R, Castellsague X, Shah KV,
Snijders PJ et al. Epidemiologic classification of human papillomavirus
types associated with cervical cancer. N Engl J Med, 2003, 348: 518-527
15 Stephen AL, Thompson CH, Tattersall MH, Cossart YE, Rose BR. Analysis
of mutations in the URR and E6/E7 oncogenes of HPV 16 cervical cancer isolates
from central China. Int J Cancer, 2000, 86: 695-701
16 Liaw KL, Hsing AW, Chen CJ, Schiffman MH, Zhang TY, Hsieh CY, Greer CE
et al. Human papillomavirus and cervical neoplasia: a case-control study
in Taiwan. Int J Cancer, 1995, 62(5): 565-571
17 Huang S, Afonina I, Miller BA, Beckmann AM. Human papillomavirus types
52 and 58 are prevalent in cervical cancers from Chinese women. Int J Cancer,
1997, 70(4): 408-411
18 Kulasingam SL, Hughes JP, Kiviat NB, Mao C, Weiss NS, Kuypers JM, Koutsky
LA. Evaluation of human papillomavirus testing in primary screening for
cervical abnormalities: Comparison of sensitivity, specificity, and frequency
of referral. JAMA, 2002, 288: 1749-1757
19 Chen X, Levine L, Kwok PY. Fluorescence polarization in homogeneous nucleic
acid analysis. Genome Res, 1999, 9: 492-498
20 Hsu TM, Chen X, Duan S, Miller RD, Kwok PY. Universal SNP genotyping
assay with fluorescence polarization detection. Biotechniques, 2001, 31:
560-568
21 Freeman BD, Buchman TG, McGrath S, Tabrizi AR, Zehnbauer BA. Template-directed
dye-terminator incorporation with fluorescence polarization detection for
analysis of single nucleotide polymorphisms implicated in sepsis. J Mol
Diagn, 2002, 4: 209-215
22 Hsu TM, Law SM, Duan S, Neri BP, Kwok PY. Genotyping single-nucleotide
polymorphisms by the invader assay with dual-color fluorescence polarization
detection. Clin Chem, 2001, 47: 1373-1377
*Corresponding author: Tel, 86-29-8541296; e-mail, [email protected]